Deep learning and digital pathology
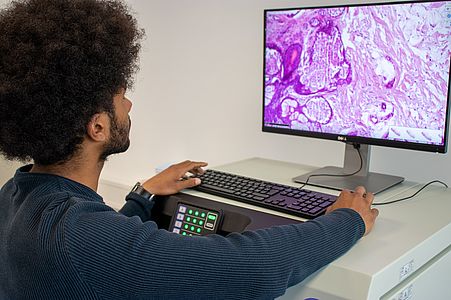
Digital pathology has recently become widely accepted due to advances in technology and regulations. Essentially, in digital pathology, a scanner creates a digital copy of the traditional glass slide that can be stored on a local or cloud-based server and processed anywhere with a computer and an internet connection. Its use for primary diagnosis has revolutionised pathology and is shaping the future of the discipline.
The DigiPath team deals with various challenges in the field of digital pathology. These include, for example, the classification of tumor and non-tumor tissue or of different tumor types based on digital microscopy images. This task would normally require extensive microscopic evaluation by pathologists. In our team, we develop powerful and robust methods for this purpose using both deep learning and classical machine learning techniques.
DigiPath-Viewer
The digipath team is also developing a "DigiPath Viewer", a web application that can be used to annotate digital microscopy images and display the results of the models. The "DigiPath Viewer" will visualize our results and also simplify the work of pathologists. Some of the main features are:
- Integration with stylus pen for natural and precise annotations.
A semi-automatic annotation tool that speeds up the annotation process. The annotation polygon can be easily adjusted by dragging/deleting points.
Visualization of automatic artificial intelligence tumor predictions as overlays on the original images. The transparency of the overlay as well as the threshold to distinguish between tumor or normal tissue can be easily adjusted.
Participation in Challenges
Our team placed 9th in the unofficial leaderboard of the CAMELYON 17 Challenge, which remained open after the official submission deadlines and received more than 100 submissions from the best companies and research groups in the field of digital pathology. The goal of this challenge is to develop a method for automated detection and classification of breast cancer metastases in whole-slide images of histological lymph node sections.
Publications
J. Le Clerc Arrastia, N. Heilenkötter, D. Otero Baguer, L. Hauberg-Lotte, T. Boskamp, S. Hetzer, N. Duschner , J. Schaller , P. Maaß.
Deeply Supervised UNet for Semantic Segmentation to Assist Dermatopathological Assessment of Basal Cell Carcinoma.
MDPI Journal of Imaging, 71 7(4), Meisenbach Verlag, Bamberg, 2021.
DOI: https://doi.org/10.3390/jimaging7040071
J. Behrmann, C. Etmann, T. Boskamp, R. Casadonte, J. Kriegsmann, P. Maaß.
Deep Learning for Tumor Classification in Imaging Mass Spectrometry.
Bioinformatics, 34(7):1215-1223, Oxford University Press, 2018.
DOI: 10.1093/bioinformatics/btx724
C. Etmann, M. Schmidt, J. Behrmann, T. Boskamp, L. Hauberg-Lotte, A. Peter, R. Casadonte, J. Kriegsmann, P. Maaß.
Deep Relevance Regularization: Interpretable and Robust Tumor Typing of Imaging Mass Spectrometry Data.
Zur Veröffentlichung eingereicht.