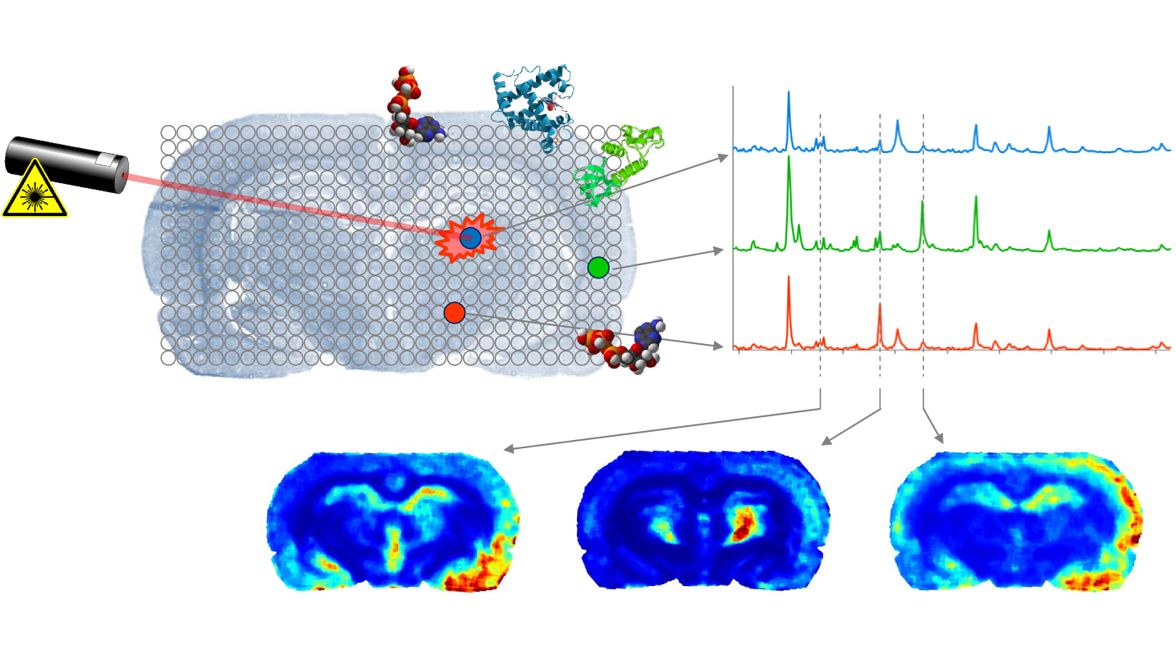
A3: MALDI imaging
Format
Block course, 1 week
Contents
Mathematical challenges of hyperspectral data analysis; theory for basic data processing methods (e.g., feature extraction, clustering and classification); importance of MALDI imaging for applications (in particular in digital pathology); hands-on work with real data.
Concepts
Mass spectrometry is one of the main data providers for genomics, proteomics or metabolomics, the related groundbreaking results in cell biology, physiology and medicine would not have been possible without appropriate tools for data analysis. Present research is now shifting from single spectra analysis towards analyzing towards spatially distributes measurements (spatial metabolomics). A key forerunner in spatial proteomics and metabolomics is MALDI imaging (matrix assisted laser desorption and ionization) mass spectrometry which is able to capture distributions of hundreds of molecules in sections of biological tissue. We start with some hands-on work (sample preparation, matrix application, measurement) in the MALDI Imaging Lab, University of Bremen. This is followed by highlighting the related bio-chemical questions, e.g., with relation to diabetes research and the quest to analyze the molecular landscape of pancreas tissue sections, see Casadonte and Caprioli (2011). The second part is devoted to numerical experiments with standard software on the acquired MALDI imaging data (data preprocessing, clustering). The major part of the course is devoted to providing the mathematical background of the basic tools used for MALDI imaging applications. We aim at including the descritpion and mathematical analysis of state of the art algorithms in MALDI Imaging based on Watrous et al. (2011).
Basic texts
- R. Casadonte, R. Caprioli. Proteomic analysis of formalin-fixated paraffin-embedded tissue by MALDI imaging mass spectrometry. Nature Protocols, 6(11):1695-1709 (2011).
- J. D. Watrous, T. Alexandrov, P. C. Doerrestein. The evolving field of imaging mass spectrometry and its impact on future biological research. Journal of Mass Spectrometry, 46(2):1521-1534 (2011).